Progeria Treated in Mice & A Shrunken Yeast Chromosome - 2021.01.11
Plus: 2D protein materials
☀️ Good morning
Omne ignotum pro magnifico est.
The Quest to Shrink Genomes
The synthetic yeast project is an international effort to build the world’s first synthetic, eukaryotic genome. Each chromosome in Saccharomyces cerevisiae is being chemically-synthesized in labs across the world. And along the way, each chromosome is receiving some bonus features, including short DNA sequences flanking each gene that enable genes to be cut out and randomly rearranged, or “scrambled”, around the genome.
For a new study, published in Genome Biology, researchers at the Shenzhen Institute of Synthetic Biology and Manchester Institute of Biotechnology “scrambled” the left arm of chromosome 12, called synXIIL, using those added DNA sequences. The chromosome was “rearranged” several times, and the team found that 39 out of 65 nonessential genes essentially dropped out after scrambling, and could be removed completely. The project is still ongoing, and the international consortium plan to meet up and put all their chromosomes together. When that happens, perhaps they could use this method to shrink down the genome en masse. Open access. Link
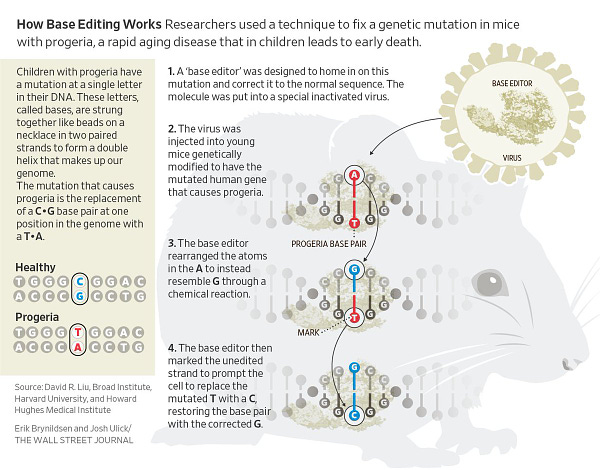
Base Editing Rescues Progeria Syndrome in Mice
Progeria is a disease that causes rapid aging. Children born with this genetic disorder—caused by a C•G-to-T•A mutation in a gene called LMNA—only live for about 14 years. For a new study, published in Nature, researchers used an adenine base editor (a protein that can convert A•T base pairs to a G•C pair instead) to correct this mutation in living mice, as well as in cultured fibroblast cells taken from children with the disease.
Delivering the base editor to the fibroblast cells resulted in an 87-91% correction of the mutation. In mice, a single injection of viruses carrying the adenine base editor extended their median lifespan from 215 to 510 days—nearly double. Link
A New Enzyme Improves Photosynthesis
Photosynthesis—the most important metabolic pathway on earth—is notoriously inefficient. RuBisCO, one of its main protein components, can fix just three carbon dioxide molecules per second. Compare that to, say, carbonic anhydrase, which can hydrate one million molecules of carbon dioxide every second.
For a new study, researchers designed and created a “new-to-nature” pathway called TaCo, or tartronyl-CoA, that can assimilate glycolate directly into central carbon metabolism. Unlike photosynthesis in the natural world, the TaCo pathway fixes carbon dioxide and does not release it. To build the pathway, the team had to create a new enzyme—one not found in nature.
In a press release from the Max Planck Institute for Terrestrial Microbiology in Marburg, Germany, a lead author, Tobias Erb, said that the TaCo pathway could be an alternative for photorespiration, but can also “be interfaced with other synthetic CO2 fixation cycles, like the CETCH cycle. Now we will be able to efficiently link synthetic CO2 fixation directly to central metabolism.” The CETCH cycle, reported in 2016, is an in vitro, synthetic pathway for carbon fixation.
Erb says that the TaCo pathway could also be used to recycle things like polyethylene terepththalate, or PET, which is found in plastics. A main component of PET is ethylene glycol, which can be directly converted to glycerate and then fed into this pathway. This work was published in Nature Catalysis, and is open access. Link

🧫 Rapid-Fire Highlights
More research & reviews worth your time
By inserting five genes into wheat crops, researchers found that the plants were protected from Puccinia graminis, a fungus that causes stem rust. The bioengineering breakthrough is important because “stem rust and leaf rust epidemics often have caused yield losses in wheat far exceeding 20 million bushels,” according to the USDA. That equates to about 1.2 billion pounds of wheat lost to rust per year in the US. Nature Biotechnology. Link
The first CRISPR-Cas systems that were discovered, y’know, out there in the wild, used two different RNAs to recognize, and cut, DNA—a crRNA and a tracrRNA. Now, researchers have found a naturally occurring single-guide RNA for S. pyogenes Cas9, which they call “long-form tracrRNA”. Cell. Link
David Baker’s lab is at it again. This time, with 2D protein materials “made by design.” For a new study, they assembled protein layers using “dihedral protein building blocks”; LEGOs anyone? Nature. Link
Citrulline is a type of amino acid that some cells “clip on” to proteins to change how those proteins behave. For a new study, researchers used an engineered leucyl tRNA synthetase-tRNA pair to incorporate citrulline into proteins at any desired position. Nature Communications (Open Access). Link
To engineer a living organism, scientists typically make one change at a time. They alter this protein, then that one, and make tweaks here and there in a somewhat hodge-podge way. A new review argues for a more statistical approach. ACS Synthetic Biology. Link
Bacterial cells use small molecules to talk to each other. Those small molecules convey a message after they bind to certain protein receptors. In a new study, researchers found that a bacteriophage (a type of virus that infects bacterial cells) called DMS3 encodes a protein that blocks one of those protein receptors, called LasR. Molecular Cell. Link
Researchers designed and built a protein sequence from scratch, and found that it could kill Mycobacterium tuberculosis, the causal agent of tuberculosis, without harming nearby lung tissue. The team behind the breakthrough calls it a “pathogen-specific antimicrobial”. Nature Biomedical Engineering. Link
To study a DNA sequence, and figure out what it encodes, scientists often mutate its letters, and then watch what happens. A new method, called Synthesis of Libraries via a dU-containing PCR Template, or SLUPT, can be used to insert mutations at multiple, specific places in a DNA sequence. OUP Synthetic Biology (Open Access). Link
Continuing with the DNA mutation thread… for a new study, researchers used a “mutating protein” to introduce up to 4 mutations per 1,000 bases of DNA, in living cells, per day. That rivals “the rate in typical in vitro mutagenesis for directed evolution of proteins,” write the authors, and means that this protein can be used for in vivo evolution of proteins. Nucleic Acids Research (Open Access). Link
For a new study, researchers implanted a “functionalized gold nanoprobe” in a living, human hepatoma cell (cancer of the liver), and show that it could be used to measure the production of hydrogen peroxide. Analyst. Link
By randomly stitching together DNA, researchers engineered yeast to produce various colors, including green, purple, and pink, by making proviolacein, violacein, and deoxyviolacein, respectively. ACS Synthetic Biology (Open Access). Link
Bacillus subtilis—a non-photosynthetic microbe—have their own 24hr circadian clocks. Science Advances (Open Access). Link
Researchers built proteins containing four different binding motifs. They then showed that these proteins could be used as “components of biosensors to monitor antibody responses and as orthogonal ligands to modulate synthetic signaling receptors in engineered mammalian cells.” Nature Chemical Biology. Link
A protein known as an epimerase—which changes the stereochemistry of molecules—was used to “convert” amino acids in proteins to non-natural variants. The researchers think it could be used to help expand the “protein alphabet”. ACS Synthetic Biology. Link
Macular degeneration is an eye disease that slowly causes vision loss, mainly in people 60 years and older. For a new study, researchers delivered an mRNA sequence encoding a Cas9 protein, as well as a guide RNA targeting a gene called Vegfa, into mice with this disease. They found that a single injection of these biomolecules into the retina “reduced the area of choroidal neovascularization by 63%”—that’s a good thing. Nature Biomedical Engineering. Link
Before a bioengineer ever gets her hands on a cell, it is useful to build a mathematical model—a framework—to predict and plan her experiments. To do that, many researchers use “whole-cell models”, a computational tool that simulates the different metabolic pathways in a cell. Unfortunately, those models don’t account for ribosomes stalling on mRNAs; a new study adds in that factor to improve these important models. Biology (Open Access). Link
Have a great week.
Until Friday,
— Niko
Bonus Tweet: David Liu explains base editors, and how they corrected a point mutation that causes progeria, or rapid aging (one of the featured papers in this newsletter).

Thanks for reading Cell Crunch, part of Bioeconomy.XYZ. If you enjoy this newsletter, please share it with a friend or colleague.
A version of these newsletters is also posted on Medium. Reach me with tips and feedback @NikoMcCarty or via email.


